Table of Contents
Orchestrated Plasticity
You will find the code to reproduce the key results of Zenke, F., Agnes, E.J., and Gerstner, W. (2015). Diverse synaptic plasticity mechanisms orchestrated to form and retrieve memories in spiking neural networks. Nat Commun 6. on https://github.com/fzenke/pub2015orchestrated
Getting the source
To run the orchestrated plasticity simulation as described in Figure 3 of the paper download or clone the pub2015orchestrated repository. Then cd into 2015orchestrated/. You need to have Auryn installed and compiled. For this code to run you either need Auryn v0.5 or if you are used to working with Auryn v0.7 or newer, use the develop branch instead which supports Auryn name spaces (introduced with version v0.7).
Running the code
Now go to the installation directory of the simulation code (when you are
reading this, chances are you are already in this directory) and update the path to your auryn installation in Makefile. Then run make.
This should build the necessary Auryn libraries that implement plasticity
and the simulation libraries (*Connection.* and sim_*.cpp). For instance the binary file sim_rc_p10c is the
one behind Figure 3 in the paper.
Finally update the output path in globalvars.sh to point to a portion of disk
with sufficient space. If you want to run distributed simulations make sure this
path is accessible from all nodes of your cluster.
Invoking ./run_orchestrated_stdp.sh on the command line
will run the three scripts 1run_init.sh, 2run_learn.sh and 3run_cued.sh in
sequence. This corresponds to the first hours corresponding to the two-fold learning and recall protocol shown in Figure 3d and j.
Output files
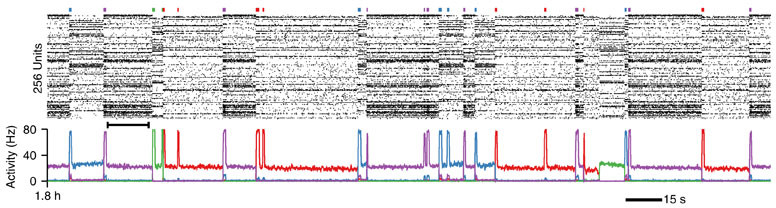
The simulation will per default run on 4 cores locally, but this might change depending on your MPI setup. It will generate a range of files out of which the most important ones are *.ras (or spk files if you use the newer develop branch version – to decode the “binary” ras files use aube). The ras files store the spike raster data of the simulation. Files labeled *.stimtimes contain the time point and duration of external stimulation events. These are text files. pact files contain the “Pattern Activity” in which patterns correspond to the cell assemblies that are formed during the course of the simulation. There is more information being saved, but these files will already allow you to reproduce Figure 3d and j (as shown above).
Phases of the simulation
Overall the code runs in three distinct and phases which are run consecutively.
The network is initialized with random recurrent weights and spatially localized input weights as described in the paper. 1run_init.sh simulates the first hour during which the network is repeatedly exposed to a barrage of four stimuli. All output files from this phase are prefixed rf1. Historic side note: rf stands for “receptive field” to distinguish this simulation from the one with random inputs which does not form stable memories in the absence of any additional homeostatic mechanism, because the random drive to network cells is too weak to push them across the LTP threshold (details are in the paper). During the first hour of simulation the actual receptive fields are formed and recurrent connections start to strengthen (cf. Figs. 3d,e in the paper). The network state is then saved.
The second phase is executed by 2run_learn.sh. It rf2 starts where rf1 stopped. Now the stimulation frequency is somewhat reduced and the network starts to exhibit delay activity after about 30-40 minutes (cf. Fig. 3j in the paper). The simulation stops after 1h total simulation time.
Finally, the simulation is continued with 3run_cued.sh and files are stored with prefix rf3. Now the network is presented with incomplete and some completely new cues to which it responds in a consistent way. This is illustrated in Fig. 4b in the paper. This simulation runs per default for another hour, but you can change the simtime value in 3run_cued.sh.
Updating rf1.pat to sort cells according to their preferred stimulus
Differences in the random seed or random library implementation might require to rebuild the rf1.pat file which stores the assembly membership of the individual neurons in the network. The file is used by PatternMonitor to read out the population firing rates of the four corresponding groups. To rebuild rf1.pat run the bootstrap.sh script. Note, that the script runs by default when you invoke ./run_orchestrated_stdp.sh.
That is also the reason why first phase of the simulation is run twice in case you wonder. After the first run the spiking data of the output is analyzed to determine cell/assembly membership relations by computing the PSTH for each cell and assigning cells to be selective to a certain pattern when their average firing rate during stimulation during the last bit of the simulation exceeds 10Hz. With the new rf1.pat generate the simulation can now rerun with the correct cells being recorded as the pattern activity (see Figure, below the spike raster).