Table of Contents
Tutorial 2: Balanced network
Here you will learn to wire up a simple balanced network. This assumes that you already know the basics explained in Tutorial 1.
The code of this example can be found here https://github.com/fzenke/auryn/blob/master/examples/sim_tutorial2.cpp
Setting up neural populations
A balanced network has an excitatory and an inhibitory population which we model as distinct NeuronGroups. Moreover, we will include some external Poisson input to jump start the network and keep activity alive.
Copy the basic Auryn simulation skeleton from Tutorial 1 to a new sim_mytute2.cpp file. In that bare bones simulation, let's first set up the network neurons:
int nb_exc_neurons = 20000; int nb_inh_neurons = nb_exc_neurons/4; IFGroup * neurons_exc = new IFGroup(nb_exc_neurons); neurons_exc->set_name("exc neurons"); neurons_exc->get_state_vector("g_nmda")->set_random(); IFGroup * neurons_inh = new IFGroup(nb_inh_neurons); neurons_inh->set_tau_mem(5e-3); neurons_inh->set_name("inh neurons");
The above code snipped initializes an excitatory population (neurons_exc) with 20000 neurons and an inhibitory population which is one quarter in size. Here were are using IFGroup one of Auryn's standard neuron models. These neurons have conductance based synapses with a fast and slow excitatory conductance. Moreover, the model has a relative refractory mechanism. Here, we initialize the NMDA conductances of the excitatory population randomly to avoid that all neurons spike synchronously initially. That's what set_random() does. Also note that we give the inhibitory neurons a membrane time constant of 5ms.
As before in Tutorial 1 we define Poisson input as a separate population:
int nb_input_neurons = 5000; float poisson_rate = 2.0; PoissonGroup * poisson = new PoissonGroup(nb_input_neurons,poisson_rate);
Connecting the network
Now let's connect these three populations. First the input:
float weight = 0.2; // conductance amplitude in units of leak conductance float sparseness = 0.05; // probability of connection SparseConnection * con_ext_exc = new SparseConnection(poisson,neurons_exc,weight,sparseness,GLUT);
And now the recurrent connctions:
float gamma = 4.0; SparseConnection * con_ee = new SparseConnection(neurons_exc,neurons_exc,weight,sparseness,GLUT); SparseConnection * con_ei = new SparseConnection(neurons_exc,neurons_inh,weight,sparseness,GLUT); SparseConnection * con_ie = new SparseConnection(neurons_inh,neurons_exc,gamma*weight,sparseness,GABA); SparseConnection * con_ii = new SparseConnection(neurons_inh,neurons_inh,gamma*weight,sparseness,GABA);
Note that we made inhibitory connections stronger by a factor of gamma = 4.0.
Set up monitors
Let's record spikes from all neurons and the membrane potential from neuron 0 in the excitatory population. To do that we set up the following monitors
SpikeMonitor * exc_spike_mon = new SpikeMonitor( neurons_exc, sys->fn("exc","ras") ); VoltageMonitor * voltage_mon = new VoltageMonitor( neurons_exc, 0, sys->fn("neuron","mem") ); voltage_mon->record_for(2);
With the last call we limit the recording time for the VoltageMonitor to 2 seconds. Because VoltageMonitor and StateMonitor have a default sampling interval of 0.1ms they quickly generate a lot of data. It's therefore advisable to limit the recording time to windows when you actually need it to speed up simulations and limit the use of disk space.
Running the simulation
To run the simulation for 10 seconds we simply add the run command:
sys->run(10);
That's it. You should now have a program which you can compile and run.
Assuming your simulation file was called sim_tutorial2.cpp and you have set up a Makefile as described here, you can simply enter this on your command line
$ make sim_tutorial2 && ./sim_tutorial
Visualizing the spikes
Running above code will generate the following files:
$ ls -ltrs | tail -n 4
912 -rwxrwxr-x 1 zenke zenke 931424 Sep 1 13:45 sim_tutorial2
4724 -rw-rw-r-- 1 zenke zenke 4835075 Sep 1 13:45 exc.0.ras
4 -rw-rw-r-- 1 zenke zenke 1077 Sep 1 13:45 sim_tutorial2.0.log
372 -rw-rw-r-- 1 zenke zenke 380022 Sep 1 13:45 neuron.0.mem
Let's take a look at the spikes (in the ras file first). In gnuplot this can be done as follows:
set xrange [2:5] set yrange [:5000] plot 'exc.0.ras' with dots lc -1
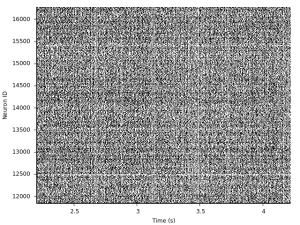 That's a lot of spikes, but the image suggests that there is some synchrony going on at times, but there are also phases of asynchronous firing. Let's zoom in a bit more:
That's a lot of spikes, but the image suggests that there is some synchrony going on at times, but there are also phases of asynchronous firing. Let's zoom in a bit more:
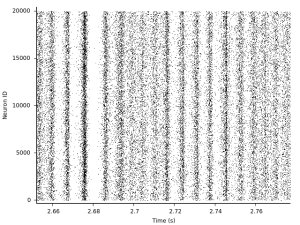
We can also analyze this a little more and plot for instance the distribution of firing rates:
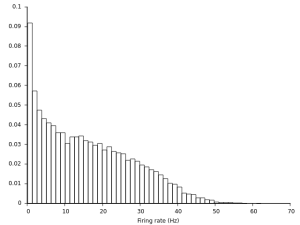
Now let's have a look the membrane trace we recorded from one of our cells:
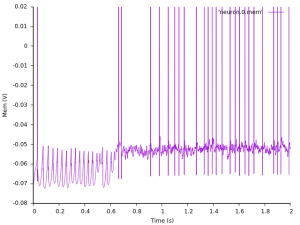 After some initial burn in period of the network dynamics the neuron seems to start firing more irregularly.
After some initial burn in period of the network dynamics the neuron seems to start firing more irregularly.
Writing a cell assembly into the E-to-E connections
Now for the fun of it, let's add a simple cell assembly to the network. Auryn support multiple ways of doing that, but the simples is to simply write a block into the weight matrix. Let's add the following code after our run instruction:
con_ee->set_block(0,500,0,500,5*weight); sys->run(2);
which increases weights in a diagonal block in the E→E connections and then runs the simulation for another 2 seconds.
Plotting again reveals the effect of this change in the spiking activity:
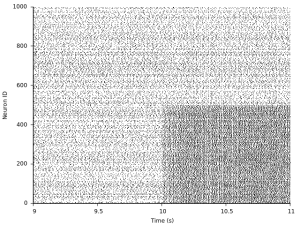
For more sophisticated ways of writing patterns or synfire chain structures into a connectivity matrix, check out the documentation of SparseConnection and specifically the functions called load_patterns (see also http://fzenke.net/auryn/doxygen/current/classauryn_1_1SparseConnection.html).
You can also always load a weight matrix from an external file which have generated using MATLAB or Python. Auryn supports a coordinate based matrix market format which allows for the seamless exchange of weight matrices with external tools (see matrix format).
In the next tutorial we will make this model plastic.
Exercises
- Run the same simulation in parallel
- Tune the connectivity parameters to asynchronous irregular firing at lower firing rates
- Add three cell assemblies and make them multi stable